Prof. Dr. rer. nat. Andreas Winterpacht
phone: +49-9131-85-22019
fax: +49-9131-85-23232
e-mail: Andreas.Winterpacht(at)uk-erlangen.de
Important publications of the last years
Spermatogenesis – single cell analyses
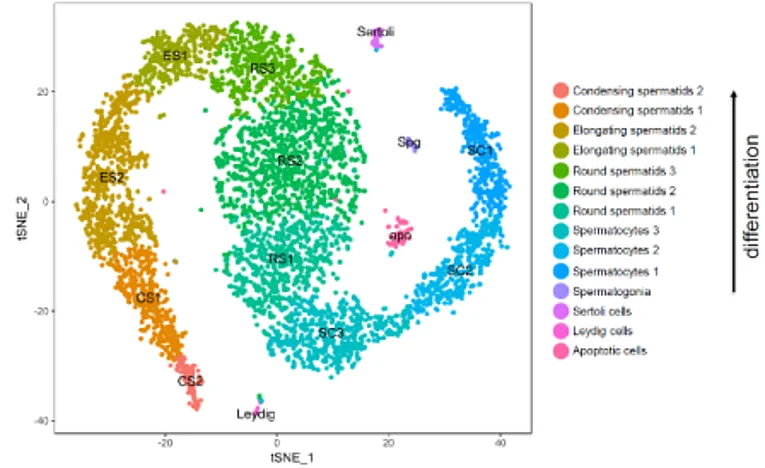
During spermatogenesis diploid spermatogonial stem cells undergo a complex and tightly regulated differentiation process to mature, haploid sperm. Proper completion of this process as well as controlling genome integrity (especially strict regulation of transposable elements) is of utmost importance for the quality and quantity of mature sperm. Defects within this process lead to male infertility which accounts for approximately half of all infertility and results from genetic abnormalities in 15-30% of cases. By employing Single Cell RNA Sequencing we were the first to describe and classify the expression profiles of the different testicular cell populations in the mouse on a single cell level. On the basis of this data, we now aim to analyze germ cell differentiation, expression/control of retrotransposons and the causes of different forms of infertility in mice and men. We focus on expression analyses, analysis of epigenetic processes and on possible transgenerational epigenetic inheritance.
The epigenetic reader PHF13
In addition, our research group is working on SPOC1 (PHF13). PHF13 codes for a protein containing a PHD-domain, two PEST-domains and a nuclear localization signal, which is expressed strongly in some rapidly proliferating immature cell types such as spermatogonia. PHF13 binds to histone methylations, thus representing an “epigenetic reader”. It is actively involved in mitosis and directly influences chromatin compaction and correct chromosome condensation during cell division. PHF13 shifts DNA double strand repair from non-homologous end joining (NHEJ) to homologous repair (HR) and is involved in co-transcriptional splicing and RNA-Polymerase II regulation.
Homozygous Phf13-/--mice are born in significantly reduced numbers, but are viable. However, they develop a progressive loss of germ cells after puberty, leading to testis hypoplasia and infertility. The underlying cause is a defect in the differentiation of spermatogonial stem cells and an increased rate of apoptosis in the pachytene stage of meiosis, where X- and Y-chromosomal transcripts are abnormally expressed. We could show that meiotic sex chromosome inactivation (MSCI) is affected. Due to the possible functions of PHF13 as an epigenetic reader and mediator of histone modifications, changes in epigenetic modifications and their interaction partners seem likely. In addition, we were able to show that PHF13 in involved in the regulation of transposable elements in the male germ line.
Thus, our research group focuses on further elucidating the role of PHF13 in the epigenetic regulation of differentiation processes as well as the regulation of retrotransposons in the male germ line.